ホーム > 尾崎 遼/ Ozaki, Haruka
尾崎 遼
Ozaki, Haruka
医学医療系 , 准教授 Faculty of Medicine

-
ヒト転写因子データの未測定範囲を体系化し研究戦略を提示
2025-09-30
尾崎 遼

-
自動実験ロボットの液体操作速度が酵母細胞に及ぼす影響を評価
2025-06-25
尾崎 遼

-
遺伝子発現の3次元分布を簡単に推計できるソフトウェアを開発
2025-01-09
尾崎 遼

-
転写因子が結合する塩基配列の新たな基盤データを構築
2023-10-12
尾崎 遼

-
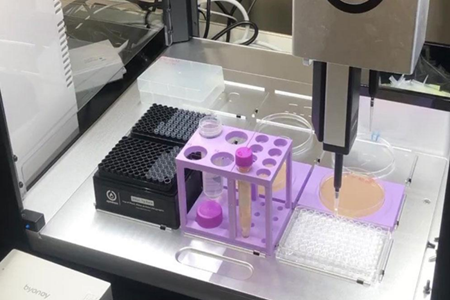
シンプルな液体分注ロボットで酵母の増殖能評価実験の自動化を実現
2023-04-12
尾崎 遼
-
細胞間相互作用による遺伝子発現を推定する解析手法を開発
2022-09-30
尾崎 遼

-
第11回生命医薬情報学連合大会 優秀口頭発表賞 生命環境学群 新井 悠也
2022-09-15
尾崎 遼

-
生命科学実験の効率的な自動化を実現するスケジューリング手法を開発
2021-08-31
尾崎 遼

-
コロナウイルスの遺伝情報に秘められた機能を解明
2020-08-25
尾崎 遼

-
見逃されていた細胞ごとのばらつきを可視化するソフトウェアを開発
2020-05-26
尾崎 遼
-
1.
Evaluation and applicability of cloud-hosted large vision-language models on the Japanese National Examination for Clinical Laboratory Technicians (Preprint)
Kota Murakami; Ryosuke Matsuzawa; Yuya Tahara-Arai; Haruka Ozaki
(2025)
-
2.
Automating Care by Self-maintainability for Full Laboratory Automation
Koji Ochiai; Yuya Tahara-Arai; Akari Kato; Kazunari Kaizu (+4 著者) Haruka Ozaki
Digital Discovery (2025)
-
3.
THG-1/TSC22D4 promotes interleukin-1 signaling through stabilization of TRAF6 in squamous cell carcinoma
Yasuhito Okano; Hiroyuki Suzuki; Yukihide Watanabe; Mohammed Abdelaziz (+14 著者) Mitsuyasu Kato
Molecular Cancer Research (2025) Semantic Scholar
-
4.
A practical guide for single-cell transcriptome data analysis in neuroscience
Yoshinori Hayakawa; Haruka Ozaki
Neuroscience Research (2025) Semantic Scholar
-
5.
Structural robustness and temporal vulnerability of the starvation-responsive metabolic network in liver of healthy and obese mice
Keigo Morita; Atsushi Hatano; Toshiya Kokaji; Hikaru Sugimoto (+15 著者) Shinya Kuroda
Science Signaling 18: (2025) Semantic Scholar
-
6.
Domain‐Shuffling in the Evolution of Cyclostomes and Gnathostomes
Hirofumi Kariyayama; Takeshi Kawashima; Hiroshi Wada; Haruka Ozaki
Journal of Experimental Zoology Part B: Molecular and Developmental Evolution (2025) Semantic Scholar
-
7.
Unmeasured human transcription factor ChIP-seq data shape functional genomics and demand strategic prioritization
Saeko Tahara; Haruka Ozaki
Briefings in Functional Genomics (2025) Semantic Scholar
-
8.
tomoseqr: A Bioconductor package for spatial reconstruction and visualization of 3D gene expression patterns based on RNA tomography
Ryosuke Matsuzawa; Daichi Kawahara; Makoto Kashima; Hiromi HirataHaruka Ozaki
PLOS ONE (2025) Semantic Scholar
-
9.
Regional developers’ community accelerates laboratory automation
Akari Kato; Takaaki Horinouchi; Haruka Ozaki; Genki N. Kanda
SLAS Technology 100211 (2024) Semantic Scholar
-
10.
Neuronal subtype-specific transcriptomic changes in the cerebral neocortex associated with sleep pressure.
Shinya Nakata; Kanako Iwasaki; Hiromasa Funato; Masashi YanagisawaHaruka Ozaki
Neuroscience research (2024) Semantic Scholar
-
11.
DNA hypomethylation characterizes genes encoding tissue-dominant functional proteins in liver and skeletal muscle
Hideki Maehara; Toshiya Kokaji; Atsushi Hatano; Yutaka Suzuki (+16 著者) Shinya Kuroda
Scientific Reports 13: (2023) Semantic Scholar
-
12.
Transcription factor-binding k-mer analysis clarifies the cell type dependency of binding specificities and cis-regulatory SNPs in humans.
Saeko Tahara; Takaho Tsuchiya; Hirotaka Matsumoto; Haruka Ozaki
BMC genomics 24: 597 (2023) Semantic Scholar
-
13.
LLMs can generate robotic scripts from goal-oriented instructions in biological laboratory automation
Takashi Inagaki; Akari Kato; Koichi Takahashi; Haruka OzakiGenki N. Kanda
(2023)
-
14.
Automation of yeast spot assays using an affordable liquid handling robot
Shodai Taguchi; Yasuyuki Suda; Kenji Irie; Haruka Ozaki
SLAS Technology (2023)
-
15.
SAGAS: Simulated annealing and greedy algorithm scheduler for laboratory automation
Yuya Arai; Ko Takahashi; Takaaki Horinouchi; Koichi TakahashiHaruka Ozaki
SLAS Technology (2023) Semantic Scholar
-
16.
Single-cell transcriptome analysis illuminating the characteristics of species-specific innate immune responses against viral infections
Hirofumi Aso; Jumpei Ito; Haruka Ozaki; Yukie Kashima (+2 著者) Kei Sato
GigaScience (2022) Semantic Scholar
-
17.
In vivo transomic analyses of glucose-responsive metabolism in skeletal muscle reveal core differences between the healthy and obese states
Toshiya Kokaji; Miki Eto; Atsushi Hatano; Katsuyuki Yugi (+19 著者) Shinya Kuroda
Scientific Reports 12: (2022) Semantic Scholar
-
18.
Identifying potential regulators of JAGGED1 expression in portal mesenchymal cells
Teppei Nishino; Masaharu Yoshihara; Takahiro Nakayama; Takaho Tsuchiya (+2 著者) Satoru Takahashi
BMC Research Notes 15: 172 (2022) Semantic Scholar
-
19.
CCPLS reveals cell-type-specific spatial dependence of transcriptomes in single cells
Takaho Tsuchiya; Hiroki Hori; Haruka Ozaki
Bioinformatics 38: 4868 (2022) Semantic Scholar
-
20.
scGR-seq: Integrated analysis of glycan and RNA in single cells.
Haruki Odaka; Haruka Ozaki; Hiroaki Tateno
STAR protocols 3: 101179 (2022) Semantic Scholar
書籍等出版物情報はまだありません。
-
1.
Millefy によるリードカバレージの細胞間不均一性の可視化
尾崎 遼
シングルセルゲノミクス研究会2019 2019年8月30日 招待有り
-
2.
シングルセルRNAシーケンス解析から考える環境DNA研究の展望
尾崎 遼
日本生態学会 第66回大会 2019年3月19日 招待有り
-
3.
Inferring activity and targets of enhancers in single cells through single-cell enhancer RNA analyses
尾崎 遼
International Workshop for Systems Genetics 2018 2018年11月19日 招待有り
-
4.
シングルセル・インフォマティクスによる細胞機能の理解
尾崎 遼
名古屋大学創薬科学研究科主催 第85回創薬科学セミナー 2018年9月25日
-
5.
(サイトメトリー研究者のための)1細胞オミクス計測データ解析のフロンティア
尾崎 遼
第28回日本サイトメトリー学会学術集会 2018年5月27日 招待有り
知財情報はまだありません。
135 total views
この研究者の他の情報源
キーワードが似ている研究者
- YING BEIWEN Bioinformatics, 遺伝子発現
- 佐野 伸行 ALUMNI Deep Learning, 深層学習
- 久米 慶太郎 Bioinformatics, 機械学習
- 土屋 貴穂 Bioinformatics, バイオインフォマティクス
- 有泉 亨 ALUMNI Bioinformatics, バイオインフォマティクス



 ORCID
ORCID
